Last year we presented results demonstrating that a deep learning system (DLS) can be trained to analyze external eye photos and predict a person’s diabetic retinal disease status and elevated glycated hemoglobin (or HbA1c, a biomarker that indicates the three-month average level of blood glucose). It was previously unknown that external eye photos contained signals for these conditions. This exciting finding suggested the potential to reduce the need for specialized equipment since such photos can be captured using smartphones and other consumer devices. Encouraged by these findings, we set out to discover what other biomarkers can be found in this imaging modality.
In “A deep learning model for novel systemic biomarkers in photos of the external eye: a retrospective study”, published in Lancet Digital Health, we show that a number of systemic biomarkers spanning several organ systems (e.g., kidney, blood, liver) can be predicted from external eye photos with an accuracy surpassing that of a baseline logistic regression model that uses only clinicodemographic variables, such as age and years with diabetes. The comparison with a clinicodemographic baseline is useful because risk for some diseases could also be assessed using a simple questionnaire, and we seek to understand if the model interpreting images is doing better. This work is in the early stages, but it has the potential to increase access to disease detection and monitoring through new non-invasive care pathways.
 |
| A model generating predictions for an external eye photo. |
To develop our model, we worked with partners at EyePACS and the Los Angeles County Department of Health Services to create a retrospective de-identified dataset of external eye photos and measurements in the form of laboratory tests and vital signs (e.g., blood pressure). We filtered down to 31 lab tests and vitals that were more commonly available in this dataset and then trained a multi-task DLS with a classification “head” for each lab and vital to predict abnormalities in these measurements.
Importantly, evaluating the performance of many abnormalities in parallel can be problematic because of a higher chance of finding a spurious and erroneous result (i.e., due to the multiple comparisons problem). To mitigate this, we first evaluated the model on a portion of our development dataset. Then, we narrowed the list down to the nine most promising prediction tasks and evaluated the model on our test datasets while correcting for multiple comparisons. Specifically, these nine tasks, their associated anatomy, and their significance for associated diseases are listed in the table below.
| Prediction task | Organ system | Significance for associated diseases | ||||||
| Albumin < 3.5 g/dL | Liver/Kidney | Indication of hypoalbuminemia, which can be due to decreased production of albumin from liver disease or increased loss of albumin from kidney disease. | ||||||
| AST > 36.0 U/L | Liver | Indication of liver disease (i.e., damage to the liver or biliary obstruction), commonly caused by viral infections, alcohol use, and obesity. | ||||||
| Calcium < 8.6 mg/dL | Bone / Mineral | Indication of hypocalcemia, which is most commonly caused by vitamin D deficiency or parathyroid disorders. | ||||||
| eGFR < 60.0 mL/min/1.73 m2 | Kidney | Indication of chronic kidney disease, most commonly due to diabetes and high blood pressure. | ||||||
| Hgb < 11.0 g/dL | Blood count | Indication of anemia which may be due to blood loss, chronic medical conditions, or poor diet. | ||||||
| Platelet < 150.0 103/µL | Blood count | Indication of thrombocytopenia, which can be due to decreased production of platelets from bone marrow disorders, such as leukemia or lymphoma, or increased destruction of platelets due to autoimmune disease or medication side effects. | ||||||
| TSH > 4.0 mU/L | Thyroid | Indication of hypothyroidism, which affects metabolism and can be caused by many different conditions. | ||||||
| Urine albumin/creatinine ratio (ACR) ≥ 300.0 mg/g | Kidney | Indication of chronic kidney disease, most commonly due to diabetes and high blood pressure. | ||||||
| WBC < 4.0 103/µL | Blood count | Indication of leukopenia which can affect the body’s ability to fight infection. |
As in our previous work, we compared our external eye model to a baseline model (a logistic regression model taking clinicodemographic variables as input) by computing the area under the receiver operator curve (AUC). The AUC ranges from 0 to 100%, with 50% indicating random performance and higher values indicating better performance. For all but one of the nine prediction tasks, our model statistically outperformed the baseline model. In terms of absolute performance, the model’s AUCs ranged from 62% to 88%. While these levels of accuracy are likely insufficient for diagnostic applications, it is in line with other initial screening tools, like mammography and pre-screening for diabetes, used to help identify individuals who may benefit from additional testing. And as a non-invasive accessible modality, taking photographs of the external eye may offer the potential to help screen and triage patients for confirmatory blood tests or other clinical follow-up.
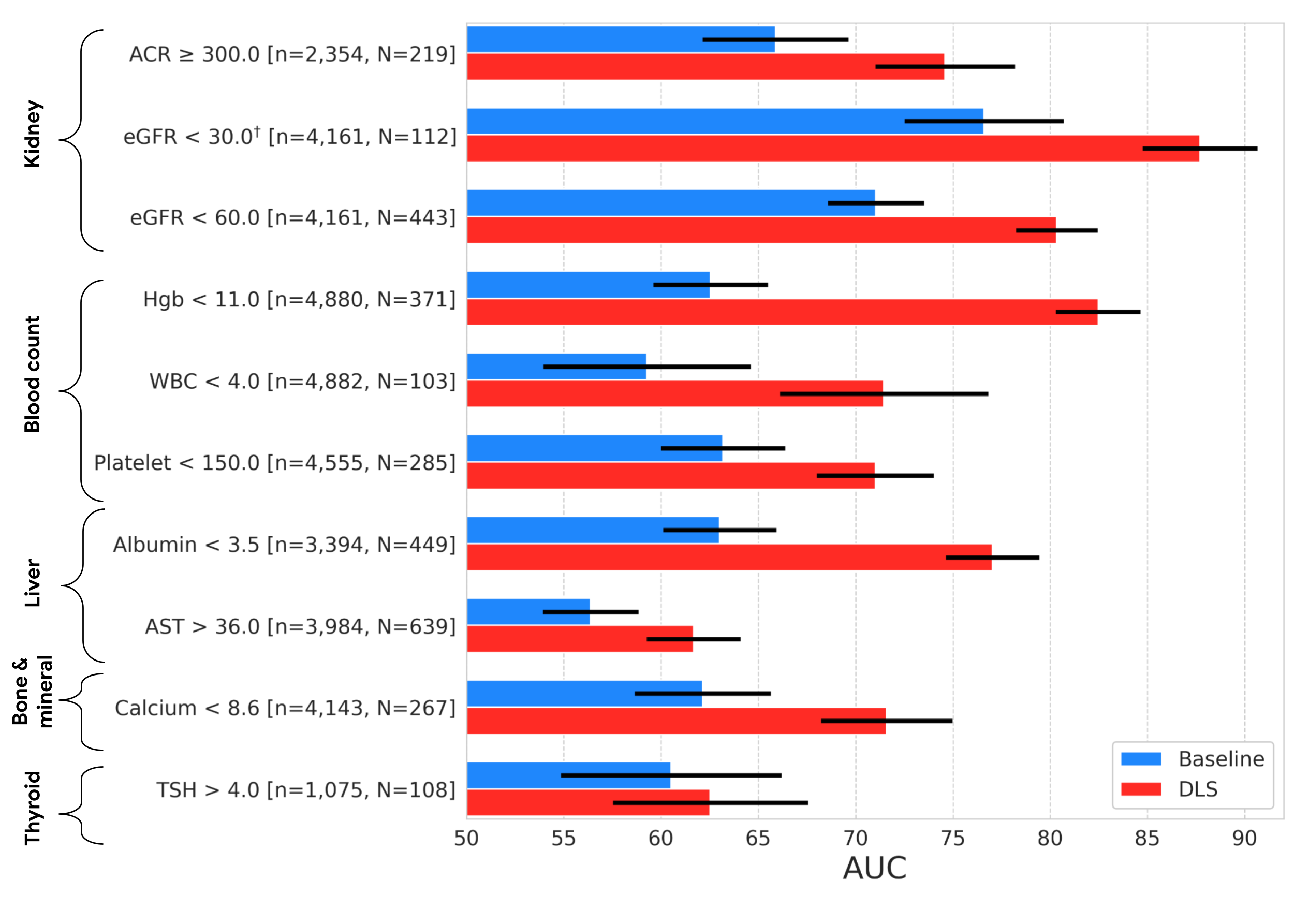 |
| Results on the EyePACS test set, showing AUC performance of our DLS compared to a baseline model. The variable “n” refers to the total number of datapoints, and “N” refers to the number of positives. Error bars show 95% confidence intervals computed using the DeLong method. †Indicates that the target was pre-specified as secondary analysis; all others were pre-specified as primary analysis. |
The external eye photos used in both this and the prior study were collected using table top cameras that include a head rest for patient stabilization and produce high quality images with good lighting. Since image quality may be worse in other settings, we wanted to explore to what extent the DLS model is robust to quality changes, starting with image resolution. Specifically, we scaled the images in the dataset down to a range of sizes, and measured performance of the DLS when retrained to handle the downsampled images.
Below we show a selection of the results of this experiment (see the paper for more complete results). These results demonstrate that the DLS is fairly robust and, in most cases, outperforms the baseline model even if the images are scaled down to 150×150 pixels. This pixel count is under 0.1 megapixels, much smaller than the typical smartphone camera.
 |
 |  |  |
| Effect of input image resolution. Top: Sample images scaled to different sizes for this experiment. Bottom: Comparison of the performance of the DLS (red) trained and evaluated on different image sizes and the baseline model (blue). Shaded regions show 95% confidence intervals computed using the DeLong method. |
Our previous research demonstrated the promise of the external eye modality. In this work, we performed a more exhaustive search to identify the possible systemic biomarkers that can be predicted from these photos. Though these results are promising, many steps remain to determine whether technology like this can help patients in the real world. In particular, as we mention above, the imagery in our studies were collected using large tabletop cameras in a setting that controlled factors such as lighting and head positioning. Furthermore, the datasets used in this work consist primarily of patients with diabetes and did not have sufficient representation of a number of important subgroups – more focused data collection for DLS refinement and evaluation on a more general population and across subgroups will be needed before considering clinical use.
We are excited to explore how these models generalize to smartphone imagery given the potential reach and scale that this enables for the technology. To this end, we are continuing to work with our co-authors at partner institutions like Chang Gung Memorial Hospital in Taiwan, Aravind Eye Hospital in India, and EyePACS in the United States to collect datasets of imagery captured on smartphones. Our early results are promising and we look forward to sharing more in the future.
This work involved the efforts of a multidisciplinary team of software engineers, researchers, clinicians and cross functional contributors. Key contributors to this project include: Boris Babenko, Ilana Traynis, Christina Chen, Preeti Singh, Akib Uddin, Jorge Cuadros, Lauren P. Daskivich, April Y. Maa, Ramasamy Kim, Eugene Yu-Chuan Kang, Yossi Matias, Greg S. Corrado, Lily Peng, Dale R. Webster, Christopher Semturs, Jonathan Krause, Avinash V Varadarajan, Naama Hammel and Yun Liu. We also thank Dave Steiner, Yuan Liu, and Michael Howell for their feedback on the manuscript; Amit Talreja for reviewing code for the paper; Elvia Figueroa and the Los Angeles County Department of Health Services Teleretinal Diabetic Retinopathy Screening program staff for data collection and program support; Andrea Limon and Nikhil Kookkiri for EyePACS data collection and support; Dr. Charles Demosthenes for extracting the data and Peter Kuzmak for getting images for the VA data. Last but not least, a special thanks to Tom Small for the animation used in this blog post.
For global asset managers, the era of AI pilots and POCs has officially passed. Today, agents are fundamentally reshaping the […]
For years, the promise of a data-driven U.S. federal government — one that can quickly and securely share information to […]
Credit: Brady Snyder / Android Authority TL;DR Google’s new compute-based Gemini limits are frustrating users who say they are hitting […]